Latest articles
Discover our most recent posts across all industries, brands and topics.
Explore our topics
Discover stories on key trends that are transforming the business and social landscape.
Company News
Keep up with the latest happenings at Dassault Systèmes.
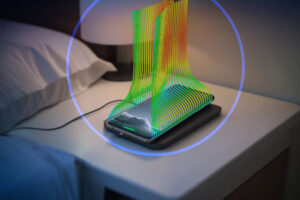
Design & Simulation
Today’s challenges are being met with innovative design and manufacturing and improve through simulation.
Editor’s picks
Explore our favorite stories
Manufacturing
It’s Fall Festival Time! Is Your Supply Chain…
Many event planners and teams focused on the supply chain may also wonder about possible…
Manufacturing
Decoding with Artificial Intelligence and Augmented Reality
New technologies offer considerable opportunities and advantages to industrial players seeking to adapt to the…
Company News
How green is your car?
The way a car’s environmental performance is measured is changing. One German engineering services provider…
Company News
How one company is leading a revolution in…
With a novel approach to reconstructive surgery, LATTICE MEDICAL is helping to shape the next…
Manufacturing
Virtual Twins for Industrial Robotics
While 34% percent of today’s biggest companies have committed to zero-emissions goals, 93% will miss…
Company News
What will healthy mean in 2040?
A new vision emerges for the convergence of life sciences innovation with global healthcare, into…
Design & Simulation
Ask an Engineer: What is Machine Learning and…
An interview with SIMULIA’s Jing Bi, a Technology Senior manager who specializes in physics-based simulation…
Latest articles from brands
Learn more about our portfolio of 3D modeling applications, simulation applications creating virtual twins, social and collaborative applications, and information intelligence applications.
Stay up to date